What is pharmacogenomics?

What is pharmacogenomics?
This blog post provides a brief overview of pharmacogenomics, beginning with the concepts of pharmacokinetics and pharmacodynamics and the definitions used in the field. It then outlines the current state of pharmacogenomics, cataloging known and actionable pharmacogenetic annotations, and concludes with examples of clinical trials investigating pharmacogenetic interactions and the challenges of conducting such studies.
March 01, 2025
Pharmacogenomics, often used interchangeably with pharmacogenetics, or short PGx, is the field that studies inter-individual variability in drug response (efficacy and/or safety) due to genetic factors. Let’s dive into some basic definitions, resources curating PGx interactions and challenges in the field.
Table of Contents
- Pharmacokinetics and pharmacodynamics
- Current state in pharmacogenomics
- Concluding remarks
- References
Pharmacokinetics and pharmacodynamics
PGx variants mostly reside in genes involved in pharmacokinetics and pharmacodynamics as well as in regions related to immune response (Lauschke et al., 2019). Pharmacokinetics can be broadly defined as what the body does to the drug and pharmacodynamics as what the drug does to the body. Pharmacokinetics evolves around absorption, distribution, metabolism and excretion (ADME) which influences the drug’s concentration and activity in the body over time. On the other hand, pharmacodynamics is concerned with the variability in drug response not related to drug concentration, but to the mechanisms of action of drugs and how they interact with specific receptors or molecular targets.
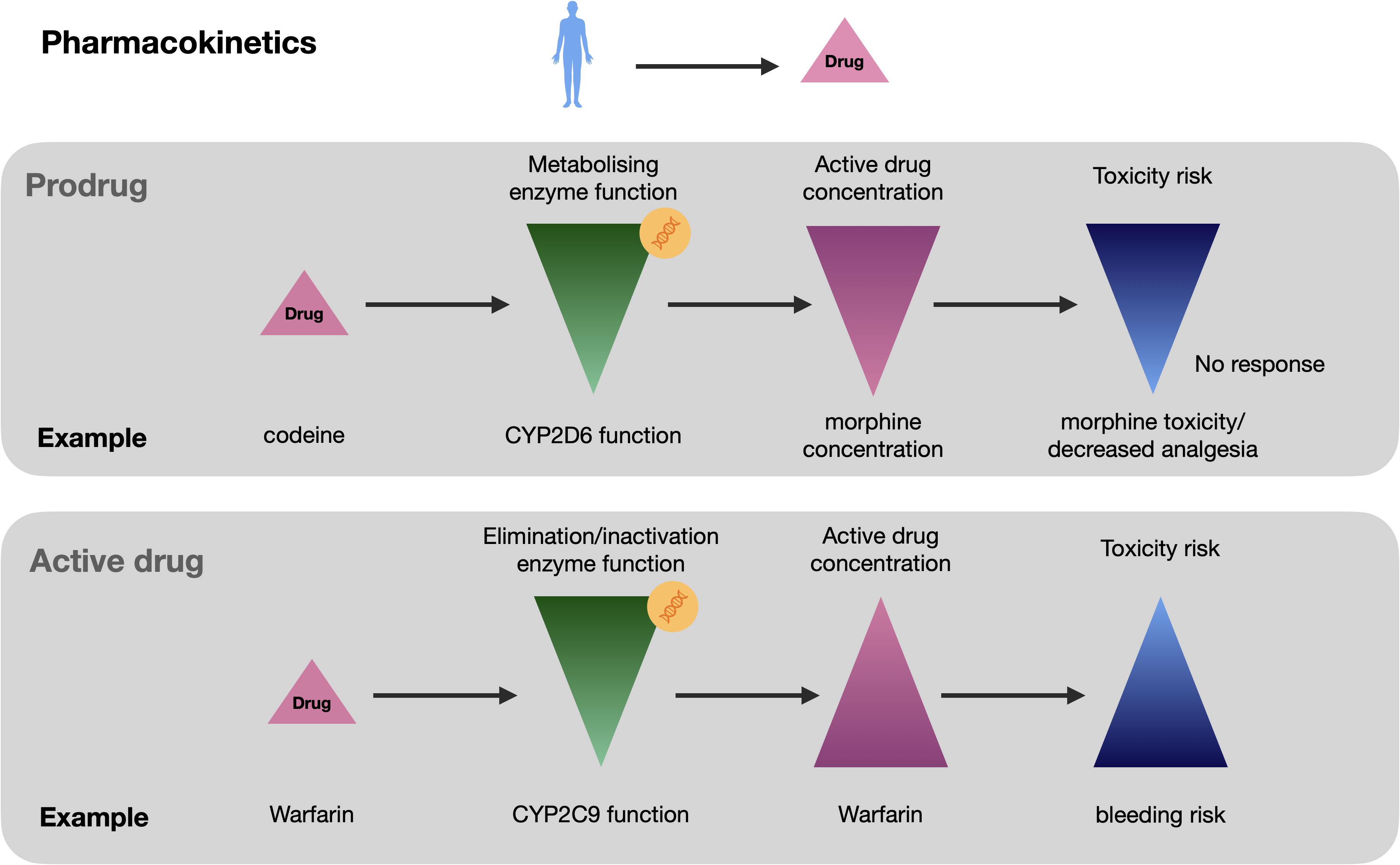
Pharmacokinetics: what the body does to the drug
Single variants in key pharmacokinetic genes can have a substantial impact on active drug concentration in two scenarios (Roden et al., 2019). In the first scenario, a prodrug needs to be bioactivated to its active drug metabolite. Variation in the responsible enzymes can lead to toxicity or lower the pharmacologic effect. Examples include the prodrug codeine that is biotransformed to morphine by the CYP2D6 enzyme (Dayer et al., 1988) as well as the antiplatelet drug clopidogrel bioactivated by CYP2C19 where LoF carriers have an increased risk of major adverse cardiovascular events (MACE) and bleeding (Mega et al., 2010). In the second scenario, variation in enzymes responsible for the elimination of active drugs with a narrow therapeutic window (i.e., small margin between therapeutic and toxic doses) can cause large PGx effects if a single pathway is responsible for inactivation or drug efflux. For instance, LoF genetic variants in CYP2C9 which is responsible for the metabolic clearance of the anticoagulant drug warfarin can increase the incidence of severe bleeding (Van Booven et al., 2010). Note that most of the clinically actionable pharmacogenetic variants described to date have a pharmacokinetic basis (Roden et al., 2019).
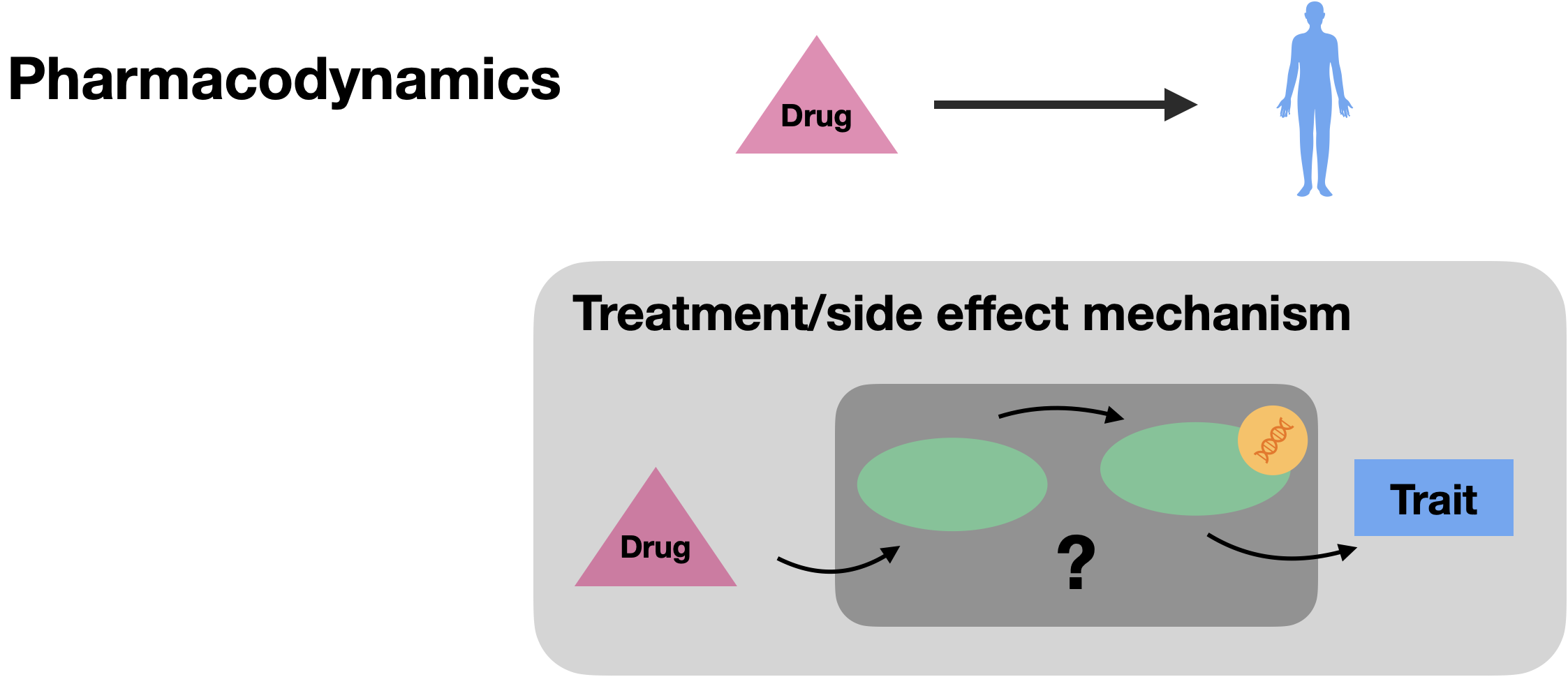
Pharmacodynamics: what the drug does to the body
Pharmacodynamic variants usually reside in genes implicated in disease mechanisms and can imply the drug target gene itself (Roden et al., 2019). For instance, variants in VKORC1, which is inhibited by warfarin, and which is involved in clotting factor activation, determine the required maintenance dose, and can also cause warfarin resistance (Rost et al., 2004). VKORC1 is one of the key pharmacogenes and the variant rs9923231C>T which is estimated to account for 15-30% of the variability is highly ancestry-specific (T allele frequency of 39%, 5% and 88% in European, African and South East Asian populations, respectively; https://www.pharmgkb.org/). Other pharmacodynamic PGx mechanisms include variants in RYR1 and CACNA1S genes that increase the risk of malignant hyperthermia upon exposure to potent volatile anaesthetics (Gonsalves et al., 2019).
Pharmacogenetics definitions
⭐ The field of pharmacogenetics is adopting a standard set of definitions to describe pharmacogenetic phenotypes
(Caudle et al., 2017).
Historically, the star allele nomenclature (*) has been used to designate pharmacogenetic alleles,
although efforts are being undertaken to harmonize the notation with alternative naming conventions such as rsIDs
(Gaedigk et al., 2018).
In the star allele nomenclature, *1 describes a fully functional (wild-type) haplotype whereas any other number refers to either a decreased or increased activity based on the presence of a single or a combination of alternative alleles. From the diplotype, a pharmacogenetic phenotype can be determined which in the case of drug-metabolising pharmacokinetic enzymes will be
normal metaboliser (corresponding to a *1/*1 diplotype or the combination of a normal and decreased function allele), intermediate, poor, rapid, or ultrarapid metabolizer.
For transporters such as SLCO1B1, which facilitates the hepatic uptake of statins, the phenotypes are increased, normal, decreased, or poor function and
for pharmacodynamic genes, phenotype definitions are positive if a high-risk allele is detected and negative otherwise
(Caudle et al., 2017).
Current state in pharmacogenomics
There are a number of resources that curate, support and expand PGx knowledge, implementation and education, a major one being ClinPGx (https://www.clinpgx.org), formerly the Pharmacogenomics Knowledgebase (PharmGKB).
ClinPGx reports clinical guideline annotations published by the Clinical Pharmacogenetics Implementation Consortium (CPIC) (Relling et al., 2011), the Dutch Pharmacogenetics Working Group (DPWG) (Swen et al., 2008), and other professional societies including the Canadian Pharmacogenomics Network for Drug Safety (CPNDS, https://cpnds.ubc.ca) and the French National Network of Pharmacogenetics (RNPGx) (Picard et al., 2017). These guidelines may have differing levels of evidence and not all are attributed to a recommendation. Also, the methodology used in scoring the level of evidence may differ between individual working groups (Abdullah-Koolmees et al., 2021). The gold standard in establishing causality between genetic variants and clinical outcomes, and demonstrating the superiority of genotype-guided treatment over the standard of care remain indisputably randomized controlled trials (RCTs). A number of large-scale RCTs have been conducted of which major trials are summarised in the following Table.
Clinical trials in pharmacogenomics
Table 1.
Major randomized controlled trials conducted to test the benefit of pharmacogenetic testing to guide treatment (click to expand)
| Name | Year (completion) | N | Drug | Gene | Endpoints | Aim | Study result | Reference |
|---|---|---|---|---|---|---|---|---|
| PREDICT-1 | 2006 | 1,956 | abacavir | HLA-B | Hypersensitivity reaction | Assessing the effectiveness of prospective HLA-B*5701 screening to prevent hypersensitivity reactions to abacavir in HIV patients | HLA-B*5701 screening reduced the risk of hypersensitivity reaction to abacavir | Mallal et al., 2008 |
| COAG | 2013 | 1,015 | warfarin | CYP2C9, VKORC1 | % of time INR in therapeutic range | Assessing whether a dosing algorithm including both clinical variables and genotype data was superior to one including only clinical variables | Genotype-guided dosing of warfarin did not improve anticoagulation control | Kimmel et al., 2013 |
| EU-PACT | 2013 | 455 | warfarin | CYP2C9, VKORC1 | % of time INR in therapeutic range | Assessing whether genotype-guided warfarin dosing was superior to standard dosing | Genotype-guided dosing was superior to standard dosing during initiation of warfarin therapy | Pirmohamed et al., 2013 |
| GUIDED | 2017 | 1,167 | antidepressants | CYP1A2, CYP2C9, CYP2C19, CYP3A4, CYP2B6, CYP2D6, HTR2A, SLC6A4 | Symptom improvement, response, and remission of depressive symptoms | Assessing whether PGx testing affects antidepressant selection and leads to better outcomes in patients with MDD | PGx testing did not significantly improve mean symptoms but improved response and remission rates | Greden et al., 2019 |
| POPular Genetics | 2019 | 2,488 | clopidogrel | CYP2C19 | MACE after PCI | Assessing the clinical utility of a genotype-guided selection of oral P2Y12 inhibitors after PCI regarding adverse events | Genotype-guided strategy resulted in lower incidence of thrombotic events and bleeding | Claassens et al., 2019 |
| TAILOR-PCI | 2019 | 5,302 | clopidogrel | CYP2C19 | MACE after PCI | Assessing the clinical utility of a genotype-guided selection of oral P2Y12 inhibitors after PCI regarding adverse events | No statistically significant reduction in MACE following genotype-guided prescription, but lower bleeding risk | Pereira et al., 2020 |
| U-PGx-PREPARE | 2020 | 6,944 | 42 drugs | 12-gene panel: CYP2B6, CYP2C9, CYP2C19, CYP2D6, CYP3A5, DPYD, F5, HLA-B, SLCO1B1, TPMT, UGT1A1, VKORC1 | Adverse reactions | Assessing the clinical utility of a pre-emptive genotyping strategy in a real-world setting | PGx-guided prescribing resulted in a 30% reduction in clinically relevant ADRs | Swen et al., 2023 |
| PRIME Care | 2021 | 1,944 | antidepressants | CYP1A, CYP2B6, CYP2C19, CYP2C9, CYP3A4, CYP2D6, UGT1A4, UGT2B1, SLC6A4, HTR2A, HLA-A, HLA-B | Proportion of prescriptions with predicted drug-gene interactions and remission of depressive symptoms | Assessing whether PGx testing affects antidepressant selection and improves outcomes in MDD | PGx testing reduced prescription of medications with predicted drug-gene interactions compared with usual care. Remission rates were not significantly higher. | Oslin et al., 2022 |
Conflicting results in clinical trials
While some of the trials showed a clear advantage of a pharmacogenetically adapted treatment strategy, other trials yielded conflicting outcomes. In the following, I list a few examples where clinical trials testing similar PGx interactions resulted in different results.
Warfarin and bleeding risk: Genotype-guided warfarin dosing resulted in a significant improvement in minimising the risk of bleeding as assessed by the international normalized ratio (INR) in the EU-PACT, but not in the COAG trial (both completed in 2013). A later trial in 2016, demonstrated the advantage of genotype-guided warfarin strategy using adverse drug reactions (ADRs) as endpoints (major bleeding, INR>4, venous thromboembolism, and death) rather than INR in 1,650 individuals.
Clopidogrel and major adverse vascular events: Patients carrying CYP2C1*2 or *3 LoF variants were found to have an increased risk of ischemic events when treated with antiplatelet medication clopidogrel, however, whereas a first trial (POPular Genetics) could demonstrate the benefits of genotype-guided selection of oral P2Y12 inhibitors after primary percutaneous coronary intervention (PCI), a second trial (TAILOR-PCI) did not.
Antidepressant medications and improvements in major depressive disorder: Conflicting results were also obtained in the GUIDED and PRIME care trials on patients with major depressive disorder (MDD), where the selection of antidepressant medication in the pharmacogenomic-guided group was adapted to have a lower potential of drug-gene interactions. The GUIDED trial found an improvement in symptoms and remission rates whereas the PRIME care trial which was longer in duration did not.
Significant reduction, but modest absolute benefit: U-PGx-PREPARE, a large-scale trial (N = 6,944) across seven European countries which tested the clinical utility of a pre-emptive genotyping strategy through a pharmacogenetic passport across 12 genes, found a 30% reduction in clinically relevant ADRs (in absolute numbers 152 (21%) of 725 in the study group with an actionable test result and 231 (28%) of 833 in the control group experienced an ADR).
Challenges in conducting PGx clinical trials
Overall, clinical trials in PGx are a challenging undertaking due to several reasons:
-
Rarity of PGx variants.
Pharmacogenetic variants often have low allele frequencies which requires large sample sizes to test for statistical significance. However, due to high costs and recruitment difficulties, this may not be feasible in practice. -
Incomplete knowledge of PGx variants.
Not all of the genetic variation determining the PGx phenotype (e.g. low metaboliser phenotype) may be known and tested. The PGx phenotype may be more polygenic than initially assumed, thus requiring a more comprehensive assessment of genetic variation to guide treatment. -
PGx knowledge is skewed toward European populations.
Relatedly, evidence of pharmacogenomic (PGx) interactions often stems from studies conducted in European-ancestry populations. An analysis of the UK Biobank showed that non-European populations carry a higher frequency of predicted deleterious variants not captured by current PGx allele definitions (McInnes et al., 2019). It has been suggested that the partial failure of the warfarin COAG trial was due in part to the large proportion of African-American participants (27%), who have a lower frequency of the tested CYP2C9 variants but may carry other untested variants influencing dosage. -
Polypharmacy and environment alter PGx effects.
Not only PGx variants, but many other factors can influence drug response among which polypharmacy. In fact, the presence of a second drug that inhibits a key enzyme can mimic the effect of a LoF variant. Although studies have been conducted to investigate such drug–drug interactions, or drug-induced phenoconversion, their scope remains limited and the real-world impact of drug-induced phenoconversion remains largely unknown. In addition to comedication, other factors such as sex, age, diet and comorbidities can influence drug metabolism and cause environment-drug-gene interactions.
Concluding remarks
Genetic variation in the population that modulates drug response has real-world implications and can lead to severe side effects. While many of these variants have been identified and catalogued using standardized definitions through large collaborative efforts, clinical implementation remains slow. Demonstrating the cost-effectiveness of PGx testing continues to be a challenge and it is one of the core objectives of the U-PGx consortium.
As drug development evolves and new modalities such as biologics become more common, it will be interesting to see whether PGx variants specific to these molecules influence treatment response. PGx variation in enzymes responsible for the activation of prodrugs or the elimination of active drug ingredients is likely specific to small molecules, but there may be variants in immune-related genes that affect the response to biologics.
Large-scale biobanks offer new opportunities to study pharmacogenetics, assess the frequency of these variants systematically across ancestries, and identify previously unknown contributors to drug response. Interesting new studies are emerging in this area, and a future blog post will explore recent research and developments in the field.
Want to read more?
For further reading, explore the resources referenced throughout this article, such as ClinPGx (https://www.clinpgx.org), as well as the other works cited here. Additionally, have a look at the review article From pharmacogenetics to pharmaco-omics: Milestones and future directions which provides a more in-depth overview of pharmacogenomics, particularly regarding the opportunities offered by large-scale biobanks (Blog Post; Publication link: https://doi.org/10.1016/j.xhgg.2022.100100 ).
References
References (click to expand)
-
Lauschke VM, Zhou Y, Ingelman-Sundberg M. Novel genetic and epigenetic factors of importance for inter-individual differences in drug disposition, response and toxicity. Pharmacology & therapeutics. 2019 May 1;197:122-52.
-
Roden DM, McLeod HL, Relling MV, Williams MS, Mensah GA, Peterson JF, Van Driest SL. Pharmacogenomics. Lancet. 2019 Aug 10;394(10197):521-532.
-
Dayer P, Desmeules J, Leemann T, Striberni R. Bioactivation of the narcotic drug codeine in human liver is mediated by the polymorphic monooxygenase catalyzing debrisoquine 4-hydroxylation (cytochrome P-450 dbl/bufI). Biochemical and biophysical research communications. 1988 Apr 15;152(1):411-6.
-
Mega JL, Simon T, Collet JP, Anderson JL, Antman EM, Bliden K, Cannon CP, Danchin N, Giusti B, Gurbel P, Horne BD. Reduced-function CYP2C19 genotype and risk of adverse clinical outcomes among patients treated with clopidogrel predominantly for PCI: a meta-analysis. Jama. 2010 Oct 27;304(16):1821-30.
-
Van Booven D, Marsh S, McLeod H, Carrillo MW, Sangkuhl K, Klein TE, Altman RB. Cytochrome P450 2C9-CYP2C9. Pharmacogenetics and genomics. 2010 Apr 1;20(4):277-81.
-
Rost S, Fregin A, Ivaskevicius V, Conzelmann E, Hörtnagel K, Pelz HJ, Lappegard K, Seifried E, Scharrer I, Tuddenham EG, Müller CR. Mutations in VKORC1 cause warfarin resistance and multiple coagulation factor deficiency type 2. Nature. 2004 Feb 5;427(6974):537-41.
-
Gonsalves SG, Dirksen RT, Sangkuhl K, Pulk R, Alvarellos M, Vo T, Hikino K, Roden D, Klein TE, Poler SM, Patel S. Clinical Pharmacogenetics Implementation Consortium (CPIC) guideline for the use of potent volatile anesthetic agents and succinylcholine in the context of RYR 1 or CACNA 1S genotypes. Clinical Pharmacology & Therapeutics. 2019 Jun;105(6):1338-44.
-
Caudle KE, Dunnenberger HM, Freimuth RR, Peterson JF, Burlison JD, Whirl-Carrillo M, Scott SA, Rehm HL, Williams MS, Klein TE, Relling MV. Standardizing terms for clinical pharmacogenetic test results: consensus terms from the Clinical Pharmacogenetics Implementation Consortium (CPIC). Genetics in Medicine. 2017 Feb 1;19(2):215-23.
-
Gaedigk A, Ingelman‐Sundberg M, Miller NA, Leeder JS, Whirl‐Carrillo M, Klein TE, PharmVar Steering Committee. The Pharmacogene Variation (PharmVar) Consortium: incorporation of the human cytochrome P450 (CYP) allele nomenclature database. Clinical Pharmacology & Therapeutics. 2018 Mar;103(3):399-401.
-
Relling MV, Klein TE. CPIC: clinical pharmacogenetics implementation consortium of the pharmacogenomics research network. Clinical Pharmacology & Therapeutics. 2011 Mar;89(3):464-7.
-
Swen JJ, Nijenhuis M, de Boer A, Grandia L, Maitland‐van der Zee AH, Mulder H, Rongen GA, Van Schaik RH, Schalekamp T, Touw DJ, van der Weide J. Pharmacogenetics: from bench to byte—an update of guidelines. Clinical Pharmacology & Therapeutics. 2011 May;89(5):662-73.
-
Picard N, Boyer JC, Etienne-Grimaldi MC, Barin-Le Guellec C, Thomas F, Loriot MA, of Pharmacogenetics FN. Pharmacogenetics-based personalized therapy: levels of evidence and recommendations from the French Network of Pharmacogenetics (RNPGx). Therapies. 2017 Apr 1;72(2):185-92.
-
Abdullah-Koolmees H, Van Keulen AM, Nijenhuis M, Deneer VH. Pharmacogenetics guidelines: overview and comparison of the DPWG, CPIC, CPNDS, and RNPGx guidelines. Frontiers in pharmacology. 2021 Jan 25;11:595219.
-
McInnes G, Lavertu A, Sangkuhl K, Klein TE, Whirl‐Carrillo M, Altman RB. Pharmacogenetics at scale: an analysis of the UK Biobank. Clinical Pharmacology & Therapeutics. 2021 Jun;109(6):1528-37.