Publications
A list of selected publications I have contributed to over the years.
Selected publications: Click on any title or image to learn more about each work.
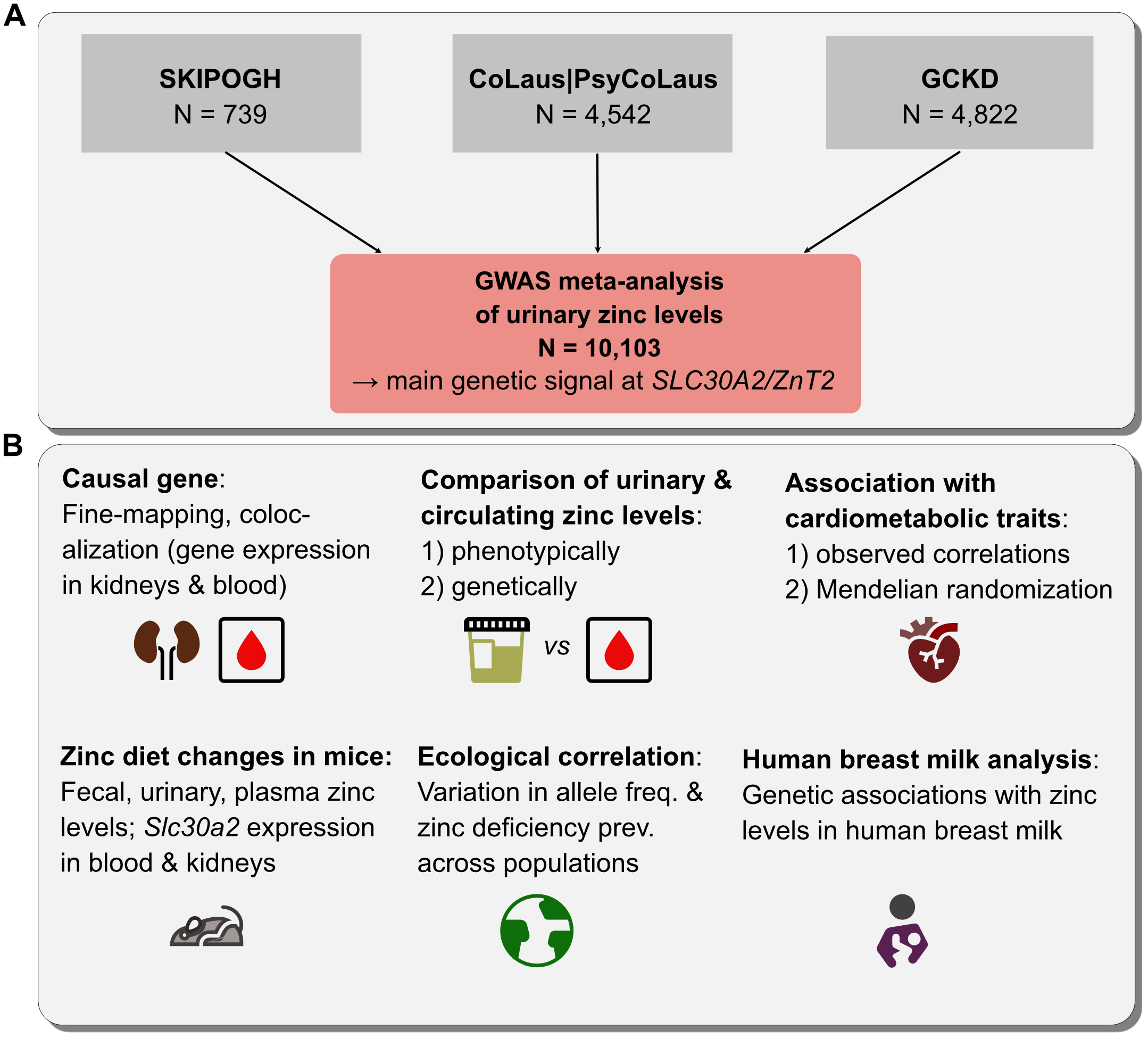
Genetic determinants of zinc homeostasis
In this study, we analyzed the genetics of zinc and its role in cardiometabolic diseases by conducting a GWAS meta-analysis on urinary zinc levels, comparing results to the genetics of circulating zinc levels and conducting follow-up experiments in mice.

Pharmacogenetics using EHRs
In this study, we analyzed electronic health records in the UK Biobank and All of Us research program to extract drug response phenotypes for ten cardiometabolic medication-biomarker pairs and conduct pharmacogenetics GWAS thereof.
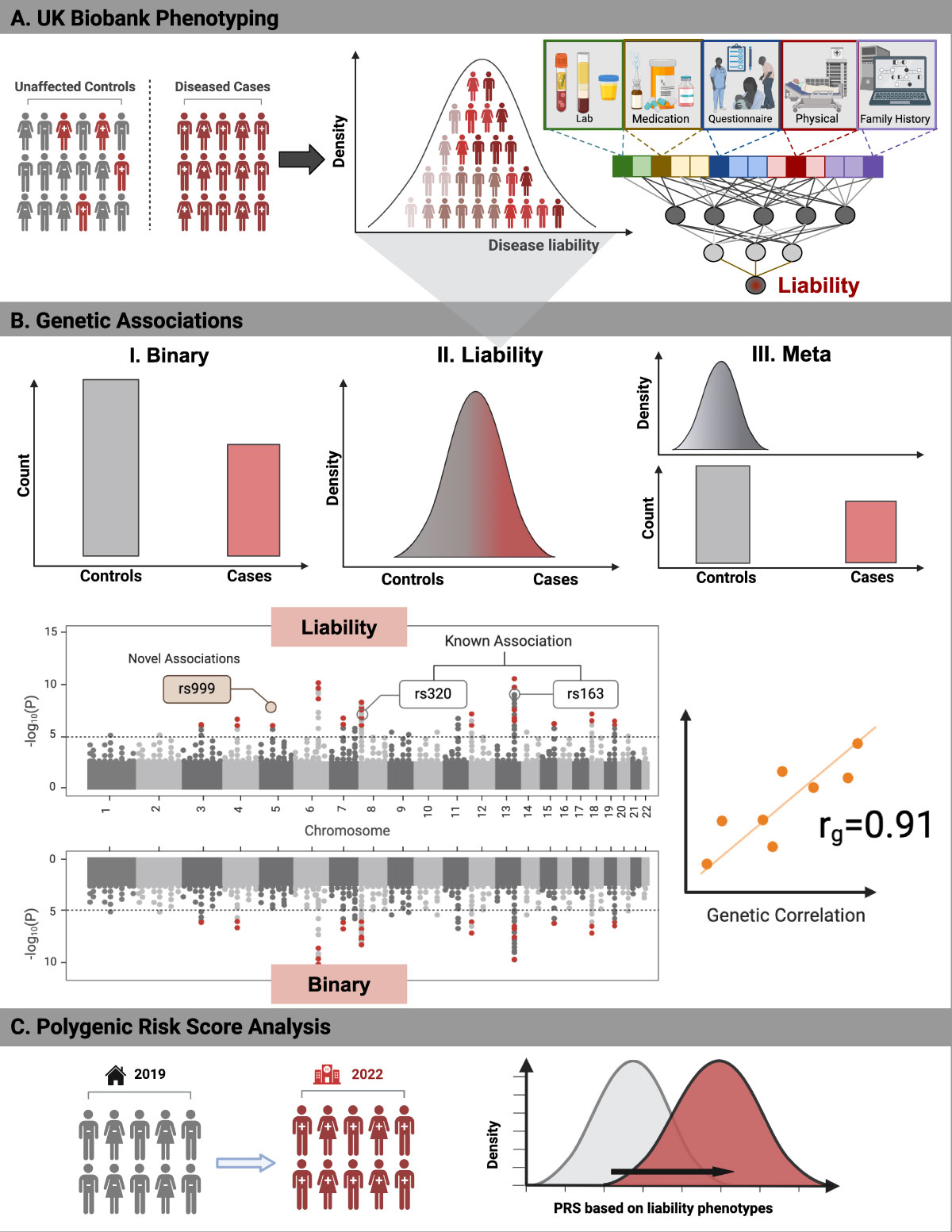
Disease liability from deep neural networks
In this study, we computed disease liabilities from deep neural networks and introduced liability and meta-GWAS methods leveraging these scores to explore genetic associations in binary traits.

Genetic support for drug targets
In this study we compared and benchmarked genetically informed approaches combined with network diffusion to prioritize drug target genes. Gene prioritization methods were based on large-scale GWAS and ExWAS, as well as tissue-wide and whole-blood expression and protein QTLs.
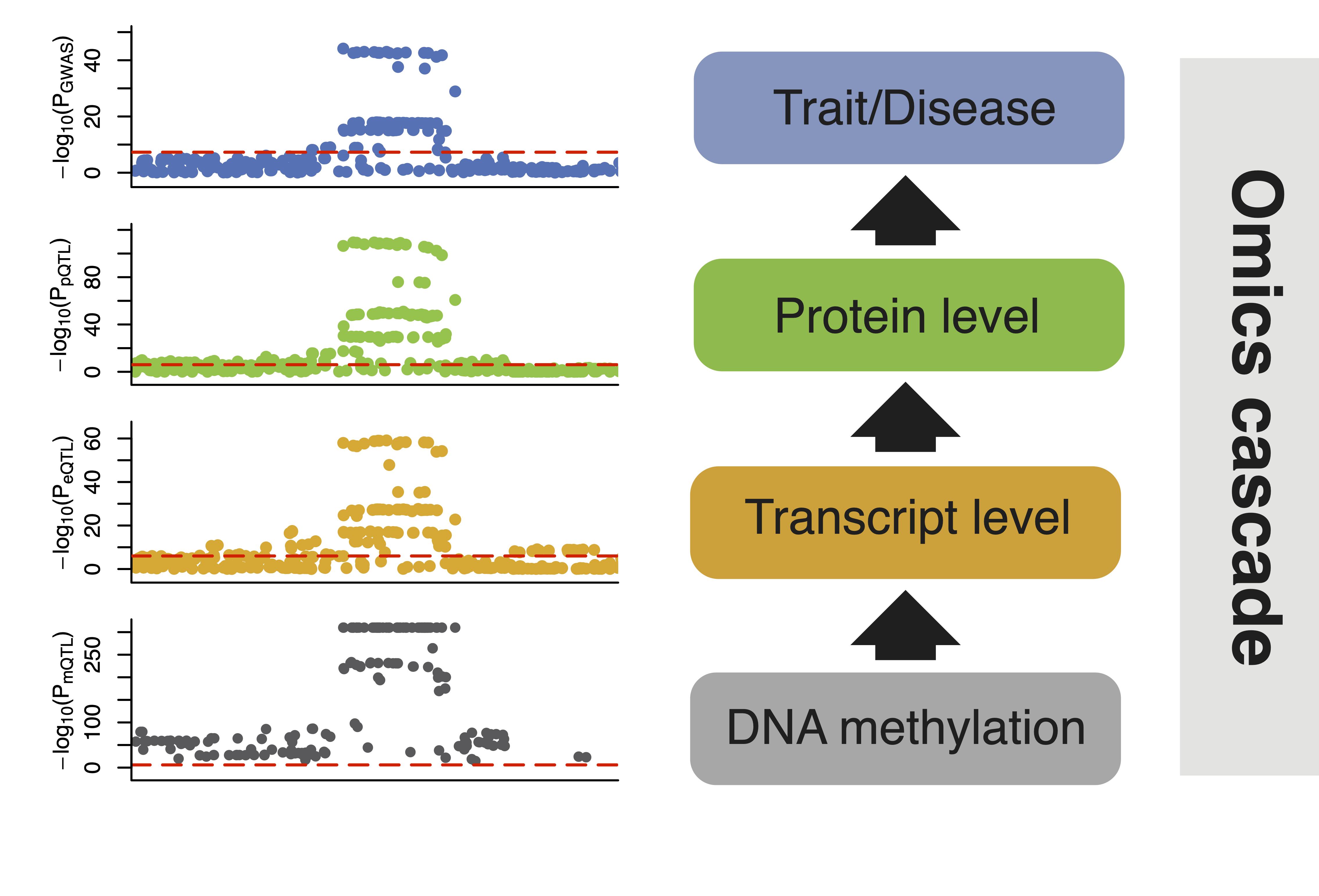
Quantifying omics mediation
We used a three-sample Mendelian randomization study design to explore the role of transcript levels in mediating DNA methylation effects on complex traits and diseases.

Review: From Pharmacogenetics to Pharmaco-omics
In this review, we discuss past, present, and future developments of pharmacogenetics methodology, with an emphasis on how multi-dimensional omics datasets can improve the mechanistic understanding of the interplay between genes and drugs.
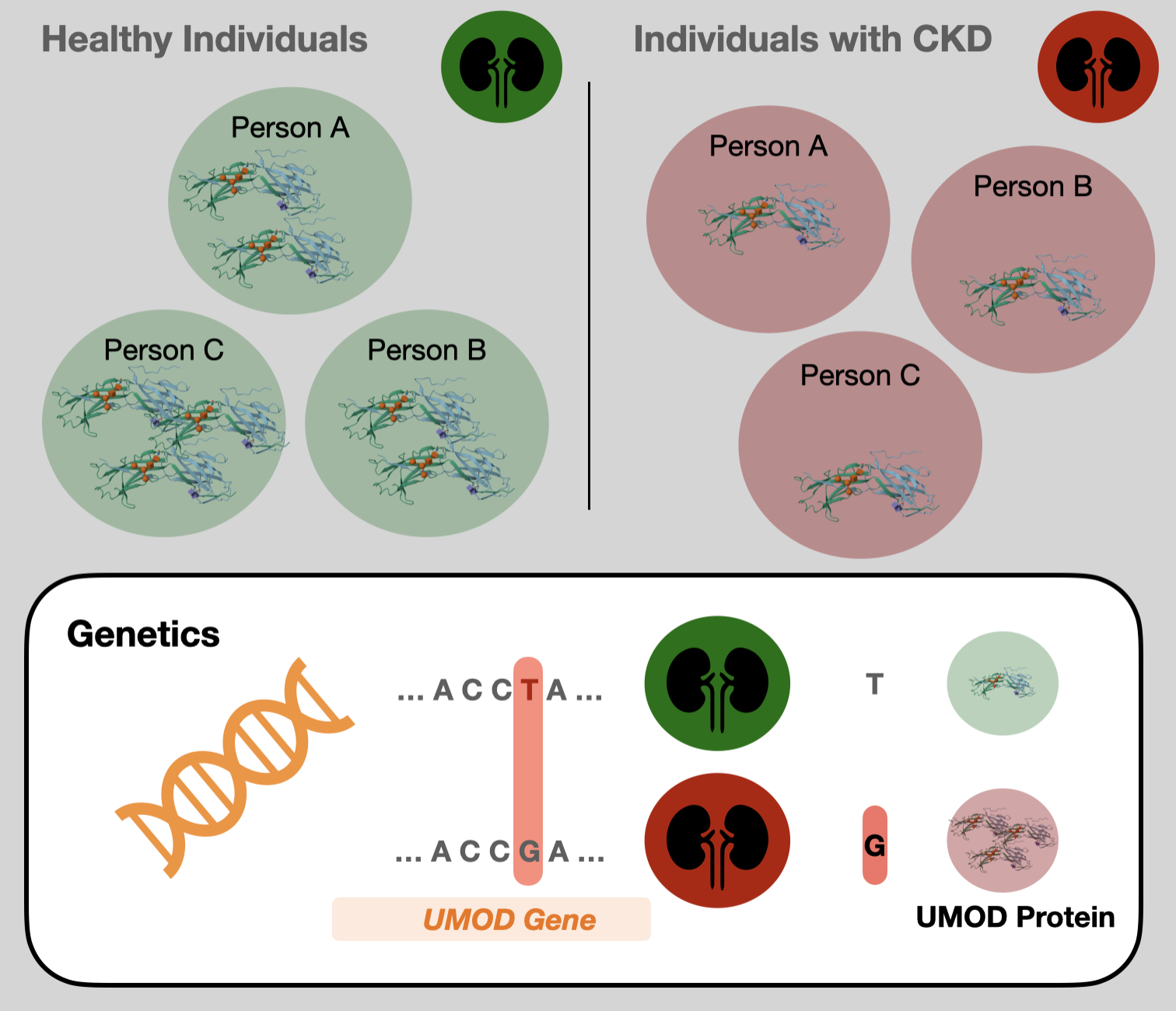
Uromodulin and chronic kidney disease
Using Mendelian randomization, we demonstrate that genetically elevated uromodulin levels have a direct, causal, and adverse effect on kidney function, opposite to the observed correlation.
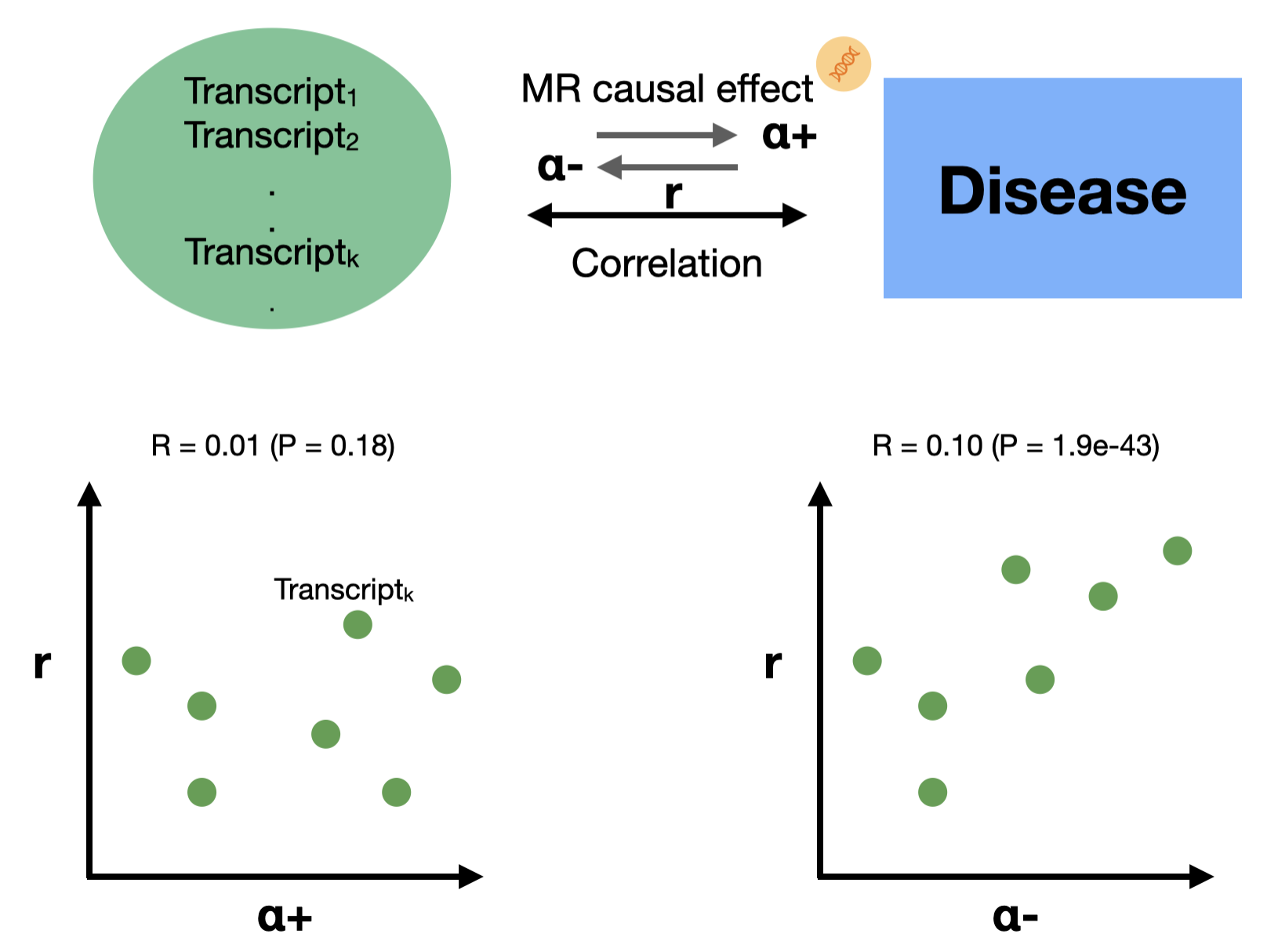
Transcriptome correlations vs causation
Using Mendelian randomization and observed transcriptomic correlations, we show that gene expression changes linked to disease more often reflect consequences of the disease than causes.
For a complete list see Google scholar